Douglas Prasher

Douglas Prasher was the first person to realize the potential of GFP as a tracer molecule. In 1987, he got the idea that sparked the GFP revolution. He thought that GFP from a jellyfish could be used to report when a protein was being made in a cell. Proteins are extremely small and cannot be seen, even under an electron microscope. However,if one could somehow link GFP to a specific protein, for example hemoglobin, one would be able to see the green fluorescence of the GFP that is attached to the hemoglobin. It would be a bit like attaching a light bulb to the hemoglobin molecule. Hemoglobin is a vital protein in the body, since it carries the oxygen in our blood. Our bodies are continuously making new hemoglobin. Encoded in the DNA is some type of index that directs the molecular machinery to the start of the hemoglobin gene. When new hemoglobin is required, protein production is activated. The gene is read and the protein is manufactured. At the end of the gene is a message called a stop codon, which ends protein production. The manufacture of proteins using the instructions from the gene is called protein expression.
Doug Prasher envisioned that it would be possible to use biomolecular techniques to insert the GFP gene at the end of the hemoglobin gene, right before the stop codon. When the cell needed to make hemoglobin, it would go to the hemoglobin gene, use the information encoded in the gene to make it, but instead of stopping when the hemoglobin was made, this cell would carry on making GFP until it reached the stop codon at the end of the GFP gene. As a result, the cell would produce a hemoglobin molecule with a GFP attached to it, see below.
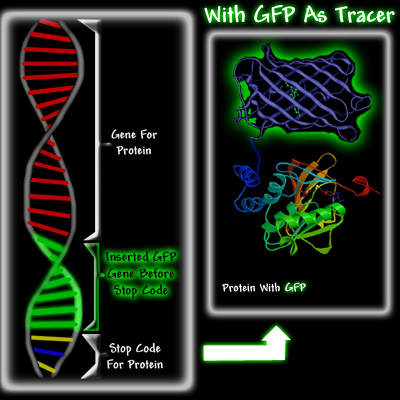
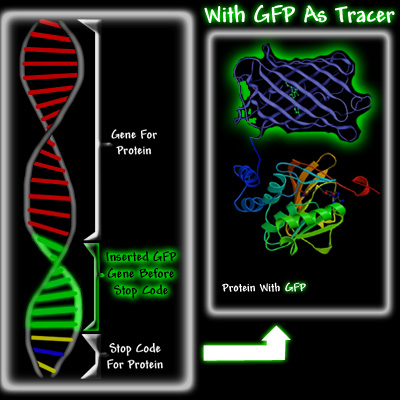
There were three reasons Prasher thought that GFP could potentially become a significant tracer molecule. Firstly, if enough protein with attached GFP was made, it should be easy to detect and to trace it as it moved through the cell, because irradiating the cell with ultra violet light would cause the GFP attached to the protein to fluoresce. Secondly, Shimomura had shown in 1974 that GFP was a fairly small protein. This was important because a small protein attached to the protein of interest was less likely to hinder its proper function. Its small size would also allow it to follow the fused protein, especially in organelles like neurons, whereas the diffusion of large proteins would be difficult. Thirdly, it had been shown that once GFP was made in the jellyfish, it was fluorescent. Most other bioluminescent molecules require the addition of other substances before they glow. For example, aequorin will glow only if calcium ions and coelenterazine have been added, and firefly luciferase requires ATP, magnesium, and luciferin before it luminesces. This would make GFP a much more versatile tracer than either aequorin or firefly luciferase, which were being used as tracers.
Besides attaching GFP to a protein and making it a fluorescent tag, Prasher also thought that GFP could potentially be a very useful reporter molecule. In order to activate protein production, DNA promoters are used. These are sequences of DNA next to genes that contain the information about where and when the gene should be read and make the protein (in other words be expressed). If GFP is linked to a specific promoter then it will be expressed in place of the protein, showing where and when the gene of interest is switched on.
Prasher’s GFP work was funded by the National Cancer Institute. In his grant he suggested that it should be possible to take the GFP gene out of the jellyfish cell and attach it to cancer cells so that they would be labeled with a fluorescent tag. Prasher managed to find the gene for GFP in Aequorea victoria and was able to express it in bacteria. In 1992, he published a paper in Gene; it reported the cloning of GFP and the sequence of the 238 amino acids in GFP, shown below. Sadly it was only a two-year grant and the funding ran out before he could express the GFP clone he had produced in a manner that would result in a fluorescent GFP.
GFP Amino Acid Sequence:
MSKGEELFTGVVPVLVELDGDVNGQKFSVSGEGEGDATYGKLTLNFICT
TGKLPVPWPTLVTTFSYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTI
FYKDDGNYKTRAEVKFEGDTLVNRIELKGIDFKEDGNILGHKMEYNYNS
HNVYIMGDKPKNGIKVNFKIRHNIKDGSVQLADHYQQNTPIGDGPVLLP
DNHYLSTQSALSKDPNEKRDHMILLEFVTAARITHGMDELYK